Resources

Scientific presentation
Making data, analysis,and AI accessible toevery scientist

Scientific presentation
Maximize AI potential in biologics discovery and development: From model training to consumption

Scientific presentation
Maximize AI potential in biologics discovery and development

Scientific presentation
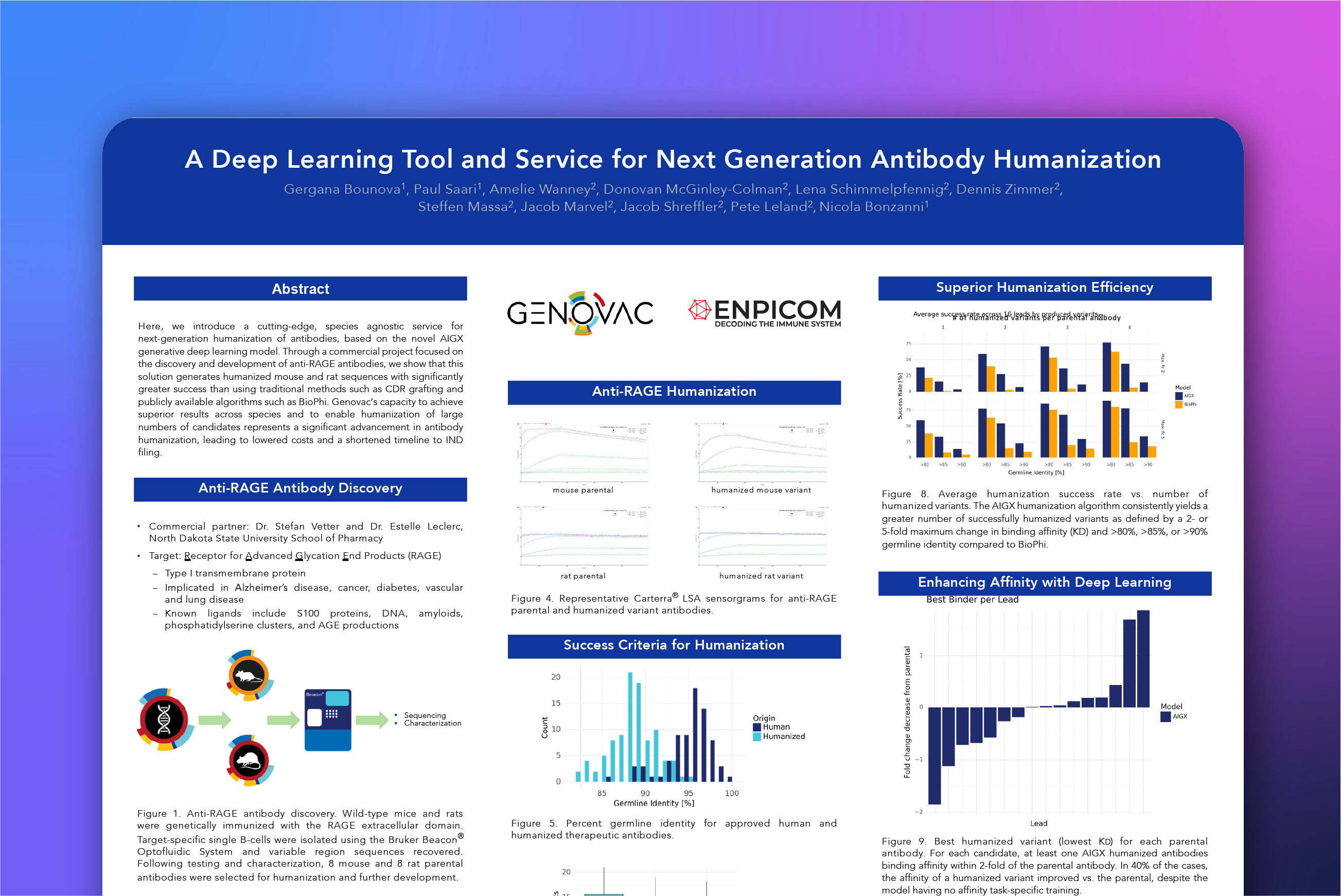
A deep learning tool and service for next-generation antibody humanization

Scientific presentation
Harnessing AI/ML in biologics discovery: Overcoming adoption challenges

Scientific presentation
Solving big data challenges in therapeutic discovery

Scientific presentation
Advances in lead discovery with high-throughput sequencing data and in-silico models

Scientific presentation
Data-driven discovery made easy, scalable, and ready for the ML era

Scientific presentation
A recipe for data-driven therapeutic discovery
Trusted by industry leaders